
Katherine L. Petrie, University of California San Diego and Justin Meyer, University of California San Diego
Evolutionary biologists like us try to figure out how organisms – from Tyrannosaurus Rex to influenza – evolve. For more than 100 years, scientists have thought about evolution through a framework that combines Mendel’s understanding of how inheritance works through genes and Darwin’s theory of natural selection that individuals better suited for their environment survive and pass on their genes to offspring.
But this framework, called the Modern Synthesis, is no longer modern; scientists came up with it in the early 20th century. Since then, biotechnology has taken off and researchers have begun to discover new phenomena that occur at the scale of molecules. These newly identified molecular processes add to our understanding of how evolution works.
One of the most frequently asked questions about the mechanism of evolution by natural selection is how it could generate new forms and functions. It’s easy to see how natural selection can improve existing functions, in a gradual, one mutation at a time manner. But it’s harder to imagine how those functions arose in the first place. As Hugo DeVries, a (very) early geneticist, put it, “Natural selection may explain the survival of the fittest, but it cannot explain the arrival of the fittest.”
In a newly published study in Science, we describe a novel mechanism for how genes evolve new functions. Our discovery is an example of how incorporating new research on molecular biology into evolutionary biology will help resolve unanswered questions.
Where do new functions come from?
How new traits come about is a big question not only because it is key to how life got from its first simple forms to the vast spectrum of biodiversity that we are a part of today, but also because answers could have immediate and practical impacts on human health. Many of the fastest evolving organisms around are pathogens; understanding why and how they evolve to evade our treatments or jump to new hosts could help us design better therapies and predict outbreaks sooner.
One way a new function might emerge is through a process called exaptation – where an existing gene is co-opted for some new function. The problem with this mechanism is that biology is rarely good at multitasking: As proteins improve at one job, they typically lose the ability to perform others.
One way nature can solve this problem is via a process called gene duplication. Through a special type of mutational event, a copy of the gene is made. Once this has happened, one copy can maintain the original version of the gene, while the other is free to accumulate mutations and take on new functions. Though there are many examples of evolution occurring by gene duplication, it was unclear if this is the only way new functions to emerge.
Birth of a new function, witnessed in the lab
To study the process of evolutionary innovation, researchers can use a technique called experimental evolution. Rather than trying to understand evolutionary events that happened in the past, experimental evolutionary biologists set up conditions in the lab where they can watch evolution happen in real time. When something interesting – like the emergence of a new function – happens, they can use all of the tools available to the modern molecular biologist to figure out how it happened.
In 2012, a team of researchers, led by one of us (Justin Meyer), was able to witness such an innovation occurring in the lab. They worked with a virus that infects the well-known bacteria E. coli. To infect their hosts, the viruses must recognize proteins present on the bacteria’s outer surface. These surface proteins are like “locks” the virus must open to enter the host cell, using its own viral protein “key.”
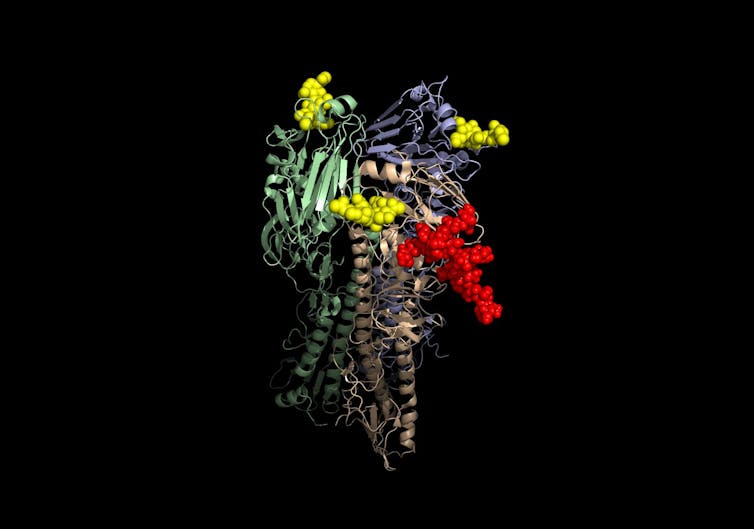
Kosik, et al. (2018), CC BY
The team set up an evolution experiment – essentially a battle royale in which the virus and the bacteria were grown together and left to fight. Over the course of three weeks and hundreds of generations, the bacteria increased security by reducing the number of “lock” proteins present on their surface. The viruses dealt with this by evolving a new version of the “key” protein that could use a different, more abundant “lock” protein as the doorway in. But surprisingly, the same virus binding protein “key” was also still able to work on the original “lock.”
How could this viral protein be good at two different functions? Genomic analysis of the evolved virus revealed that it did not undergo a gene duplication. Rather it had acquired four mutations in the genetic code that caused changes in the amino acids sequence of the resulting protein. We reasoned that these mutations must somehow make the virus able to use two different types of bacterial “locks,” however the mechanism for how these mutations worked remained unknown.
One genotype, two phenotypes
The crucial step we took in our new research was to imagine that a core idea of molecular biology, that one gene produces one protein, could be wrong. By opening our minds, now, we were able to figure out the hidden mechanism for how the mutations worked.
It turns out that the mutant protein chain the virus produces is capable of folding into different structures. Depending on chance processes during folding, sometimes it folds into a protein that can bind the old receptor, and sometimes it folds into a protein that can bind the new.
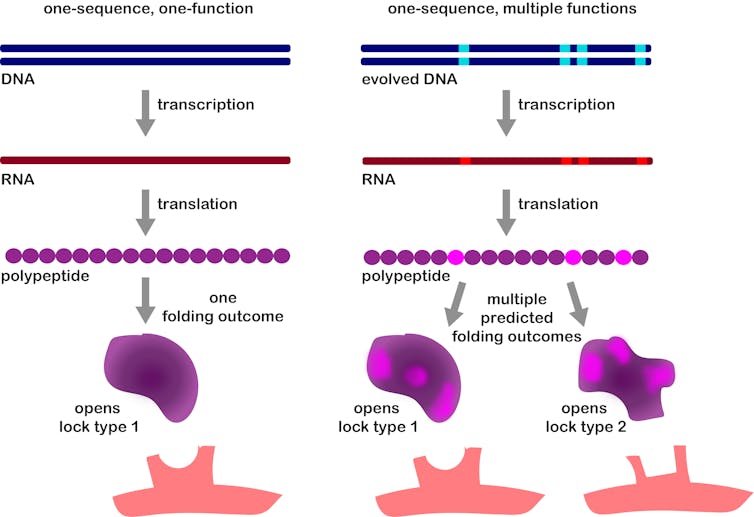
Katherine L. Petrie, CC BY-ND
Every newborn virus has about a 50-50 chance of having one version of the protein or the other. This structural variation can’t be passed on: When the newborn viruses themselves reproduce, each of their progeny viruses also has a 50-50 chance for having the new function. Since every virus produces multiple progeny, in every generation there will be viruses that can bind both receptors.
Once this intermediate version of the gene – a gene that was actually good at multitasking – was established, then conventional natural selection could take over. When we grew the viruses with bacteria that only produce the new receptor, natural selection was able to optimize the gene to specialize on it, and a more stable version of the gene to make the desired “key” protein emerged.
Nongenetic variation and evolution
This innovation is due to what scientists call nongenetic variation. A single gene sequence makes both proteins, but variation occurs as a result of chance during the 3-D folding processes when the proteins are produced in the cell. In other words, even with an identical gene sequence in all the viruses, the resulting protein varies randomly.
Evolutionary biologists have long argued over the potential role of nongenetic variation in fostering evolutionary innovation. Now we’ve seen a mechanism that takes advantage of it in action. For the virus, the nongenetic variation in protein structure is a way to get more functional bang for its genomic buck – it gets two functions for the price of one and avoids the pitfalls of trying to optimize two functions at once.
We’ve discovered a new, possibly general, mechanism for evolution that helps explain how adaptation can happen so fast. More broadly, it shows how messiness in biology – in this case an imperfect sequence-function relationship between gene and protein – can actually be an opportunity to exploit, rather than a problem to be solved.
![]() The prebiotic chemist Leslie Orgel was famous for his maxim, “Evolution is cleverer than you.” Experimental evolution and modern techniques demonstrate just what he means: When we watch evolution happen, we might be surprised at what it reveals.
The prebiotic chemist Leslie Orgel was famous for his maxim, “Evolution is cleverer than you.” Experimental evolution and modern techniques demonstrate just what he means: When we watch evolution happen, we might be surprised at what it reveals.
Katherine L. Petrie, Assistant Teaching Professor, University of California San Diego and Justin Meyer, Assistant Professor of Biological Sciences, University of California San Diego
This article was originally published on The Conversation. Read the original article.




















